Comprehensive Antibiotic Resistance Database (CARD)
Tech ID
15-055
Inventors
G. Wright
A. McArthur
Website
Commercial Enquiries
Contact
Rimika Sachdeva
Business Development
Overview
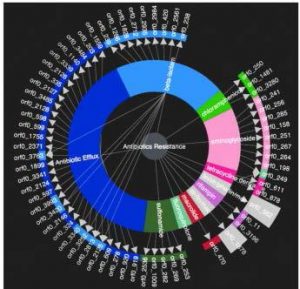
The CARD is an ontology- and model based framework for detection of antibiotic resistance genes. It contains an expert-curated collection of characterized, peer-reviewed resistance determinants and associated antibiotics, organized by the Antibiotic Resistance Ontology (ARO) and AMR gene detection models. The software utilizes these detection models for predicting AMR from genome sequences.
Other associated tools include the Resistance Gene Identifier (RGI) which is a novel genome analysis tool for resistome prediction. It annotates DNA and protein sequences and detects matches to CARD reference sequences, variants of known AMR genes and emergent threats.
Resistomes is a computer-generated data set including genome annotation and variants data using RGI for 85 pathogens of interest. For each of these pathogens, complete chromosome sequences, plasmid sequences, and whole genome shotgun assemblies were analyzed individually by RGI to generate prevalence data. Results are further categorized using the ARO.
Benefits
- Expert-curated database that is updated regularly.
- As of Feb 14, 2024: This update includes addition of gyrA, gyrB, and folp mutations for additional Salmonella species, 23S rRNA mutations for Treponema pallidum, and gyrA mutations for Helicobacter pylori. It also includes addition of OXA beta-lactamase sub-families, with subsequent re-organization of the OXA beta-lactamase classification.
- As of Oct 10, 2023: This update includes improved curation of 16S rRNA mutations, the addition of TEM and OXA beta-lactamase sequences, improvements in tet gene curation, additional aminoglycoside resistance genes, and improvements to antibiotic and drug class curation. In addition to the database, the Resistance Gene Identifier software has been updated to use BLAST 2.14.0, with config file, and adds HSP subject coordinates and impacted antibiotics to output. https://github.com/arpcard/rgi
- As of June 23, 2023: The in silico Resistomes, Variants, & Prevalence data set has been updated to 381 pathogens, 24291 chromosomes, 2662 genomic islands, 48212 plasmids, 172216 WGS assemblies, 276270 alleles based on sequence data acquired from NCBI on May 2, 2023 and Islandviewer 4, analyzed using RGI 6.0.2 (DIAMOND homolog detection) and CARD 3.2.7. Includes pre-compiled k-mer classifier data for pathogen-of-origin prediction.
- As of May 26, 2023: CARD curation updates include continued improvements for aminoglycoside modifying enzymes, curation of Mycobacterium tuberculosis mutations, and curation of additional TEM, EBR, IND, KPC, LAQ, NDM, WUS, and IMP beta-lactamases.In addition, the CARD team are happy to announce that the Resistance Gene Identifier (RGI) has been incorporated into the Chan Zuckerberg ID platform for free, cloud-based metagenomics analysis by researchers, allowing high-throughput use of RGI’s read-mapping algorithms online: https://chanzuckerberg.zendesk.com/hc/en-us/articles/15091031482644-AMR-Pipeline-Workflow
- Software for prediction of resistance genes from isolates or metagenomic samples from clinical, environmental, or agricultural samples
Applications
- Predict antimicrobial resistance (AMR) from genome sequences
- Metagenomic analysis
- Pathogen of origin prediction
- Prevalence of AMR determinants
- Regulatory submissions and product registration for agricultural products
- Commercial services to analyze genomes
- Understanding molecular mechanisms of drug resistance, particularly drug development
Copyright
Unrestricted research use of CARD and all associated data and tools is available to non-profit organizations, governments or universities. All commercial entities must seek to obtain a non-exclusive license by filling out the License Request Form.
References
Alcock, P., Huynh, W., Chalil, R. et al. CARD 2023: expanded curation, support for machine learning, and resistome prediction at the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 2023 Jan 6;51(D1):D690-D699. doi: 10.1093/nar/gkac920.

